Unique Info About How To Detect Snp

Find your snps on the chromosome.
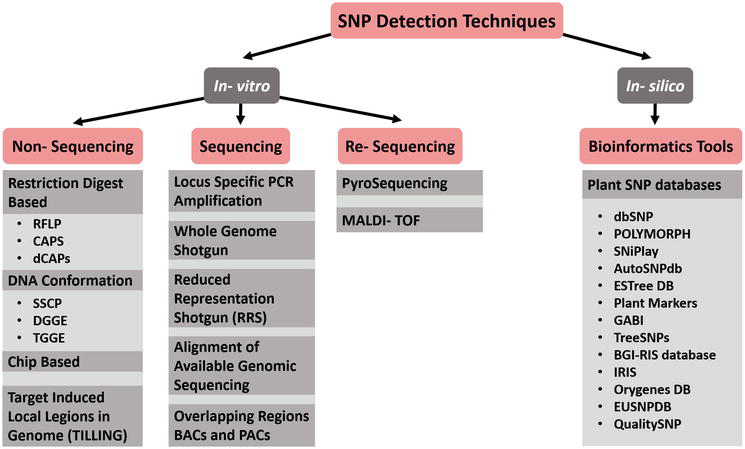
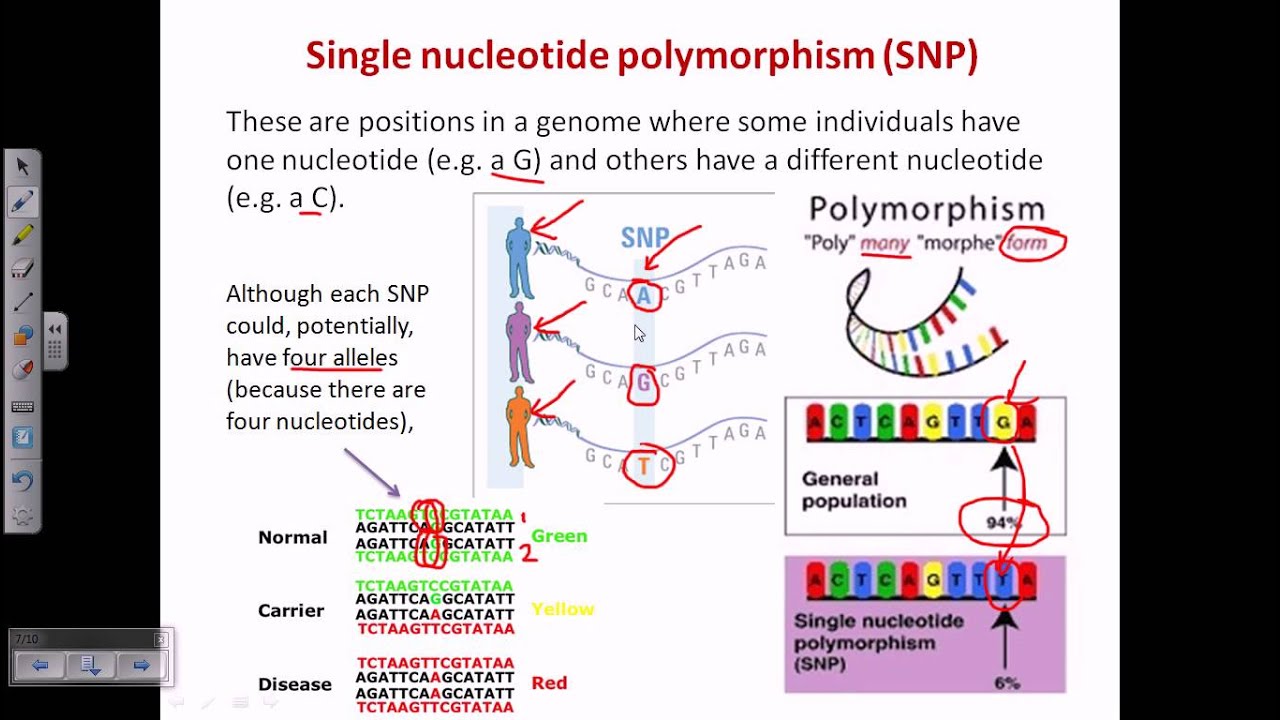
How to detect snp. What is the best way to detect snps? Their characteristics, how snp markers are developed? In testing the performance of mrsnp on snps experimentally validated to affect mirna binding, mrsnp correctly identified 69% (11/16) of the snps disrupting binding.
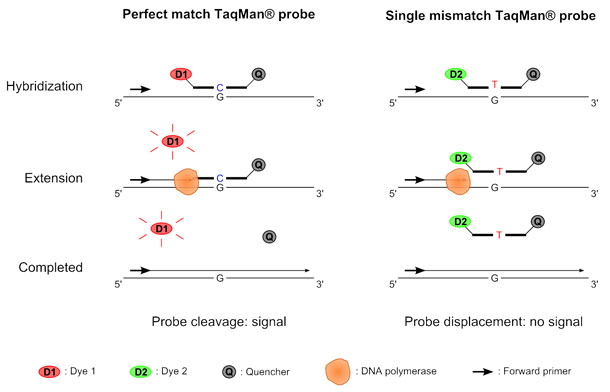
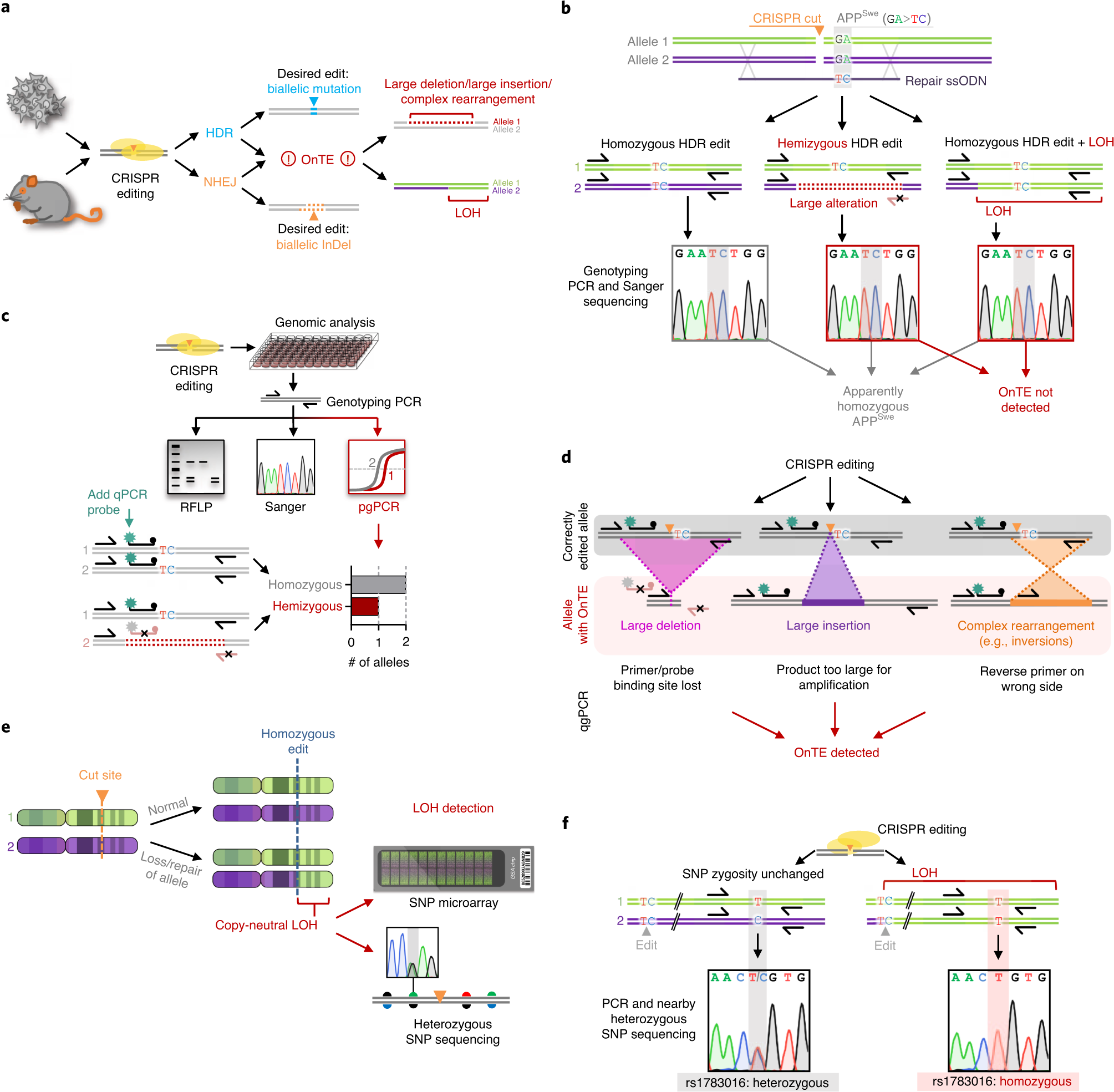
Samples are then analyzed using illumina bluefuse software and compared to a. Unlike gene expression qpcr, snp detection requires two probes with different fluorescent reporters. Snp microarray uses known nucleotide sequences as probes to hybridize with the tested dna sequences, allowing qualitative and quantitative snp analysis through signal.
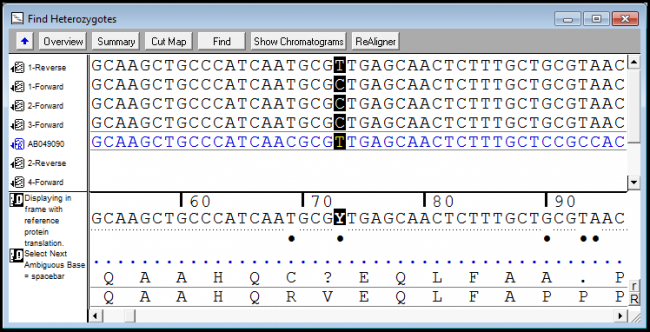
Next you will need to go to a website called ensembl. What are snp markers, why they are so popular? Sequencher has several powerful tools to help you detect mutations and snps in your dna sequences.
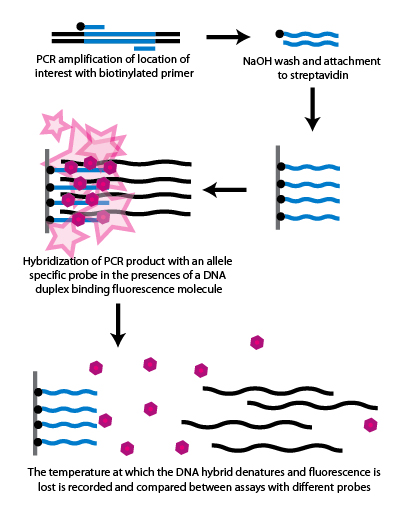
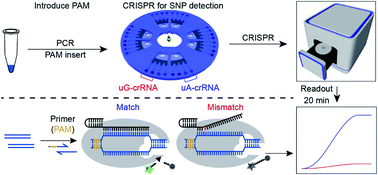
Such as, inserting sg into dna double strands, and then immobilizing dna on the surface of. Any information about the time required and about the costs of each one is also. Restriction fragment length polymorphism (rflp) is considered to be the simplest and earliest method to detect snps.
Wicell currently utilizes the cytosnp 850k microarray from illumina (illumina™ infinium snp microarrays). If possible, avoid placing an lna nucleotide at the extreme 3’ end of the primer, as this can result in low. On the homepage there is a dropdown box where you can select your organism of.
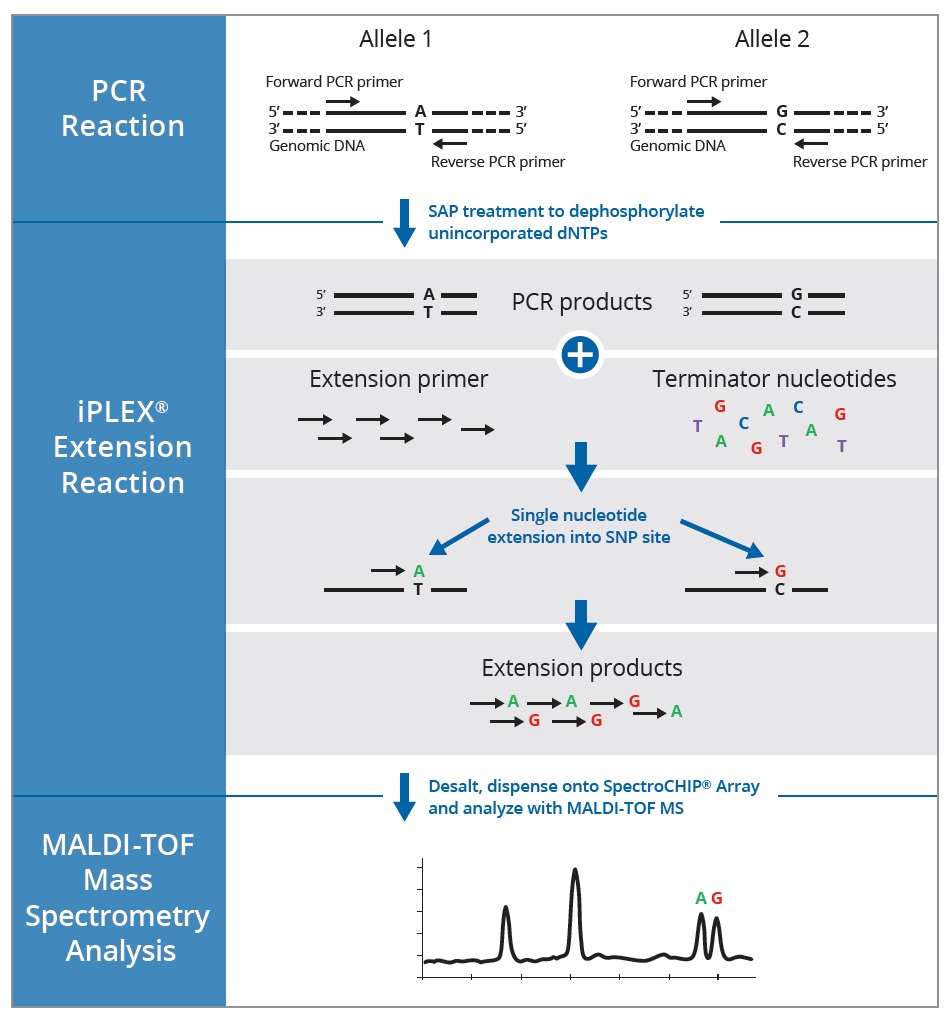
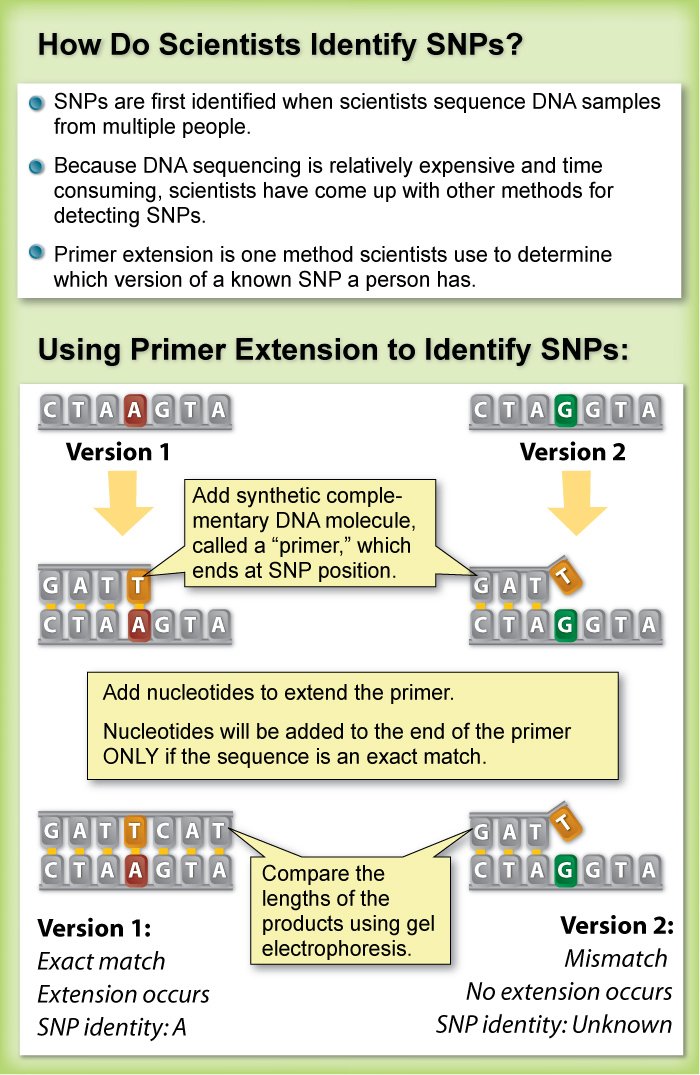
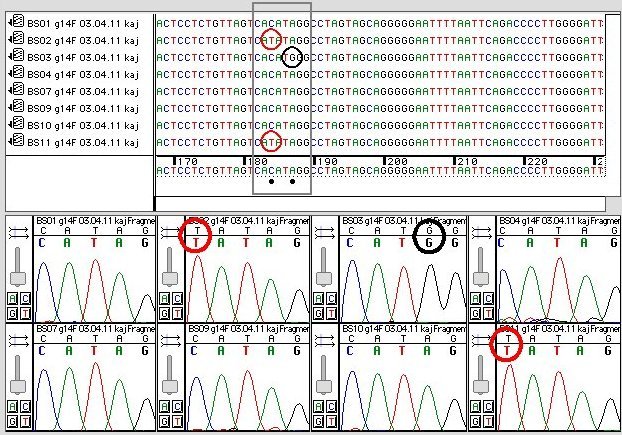
This allows us to differentiate homozygous from heterozygous samples [5]. Thus, a fast, economical and reliable sg embedded sensor is introduced to detect snp. Methods of snp detection will be covered in this video.*.
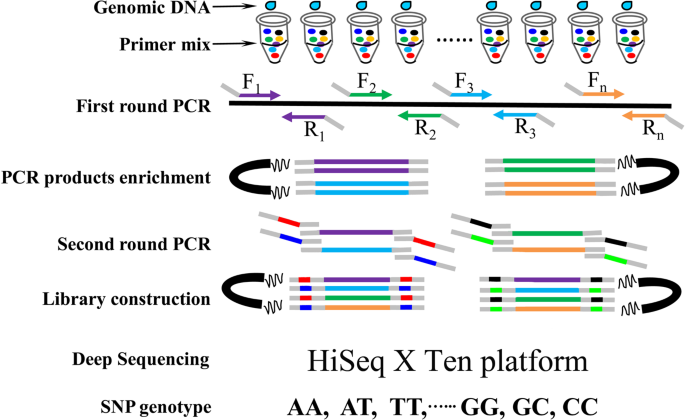
We have designed a snp identification pipeline.